Laboratory of Cell Network Regulation
1. Members
| Project leader | OKADA Mariko |
|---|
2. Background and objectives
The signal transduction network is a system that connects the extracellular environment and genetic information, and these abnormalities are known to be involved in the development of human diseases. The signal transduction system has the property of changing the difference in affinity between ligand and receptor binding to the difference in cell fate. In a system with high non-linearity where a slight difference in input occurs as a large difference, not only experimental approach but also an approach using computation and mathematical models is required. The purpose of this laboratory is to clarify the regularity and control mechanism of signal transduction and transcriptional regulation networks in human diseases such as cancer and inflammation, and to utilize that knowledge for cell manipulation and drug identification.
3. Overview of our research
3-1.Mathematical modeling of signaling networks
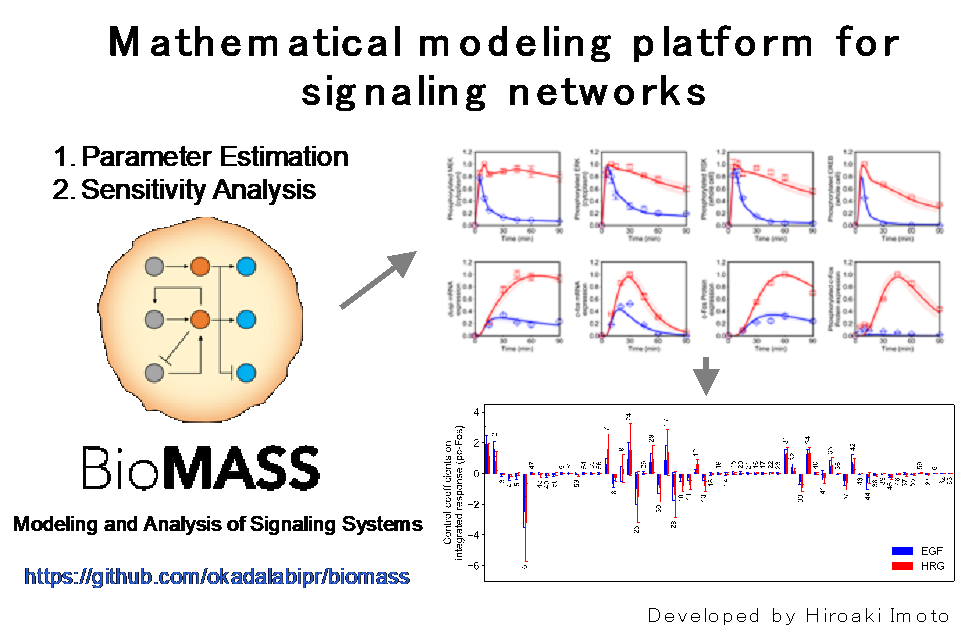
Cells make fate choices such as proliferation, differentiation, and cell death according to the external environment. Environmental information is processed by the signal transduction pathways, but since there are many types of signal transduction pathways and they have crosstalk and feedback control, the relationship between input and output is not well predicted. These properties may make it difficult to quantitatively understand drug responses to signaling network. In order to understand the characteristics of such a system, we are using mathematical modeling. We have constructed mathematical models such as the ErbB receptor-ERK-AKT pathway in cancer and the NFkB pathway in inflammation. We make these models publicly available and developed BioMASS (Modeling and Analysis of Signaling Systems) platform for these simulation analysis. The BioMASS package can be downloaded for free from https://pepy.tech/project/biomass.
3-2.Patient-specific modeling
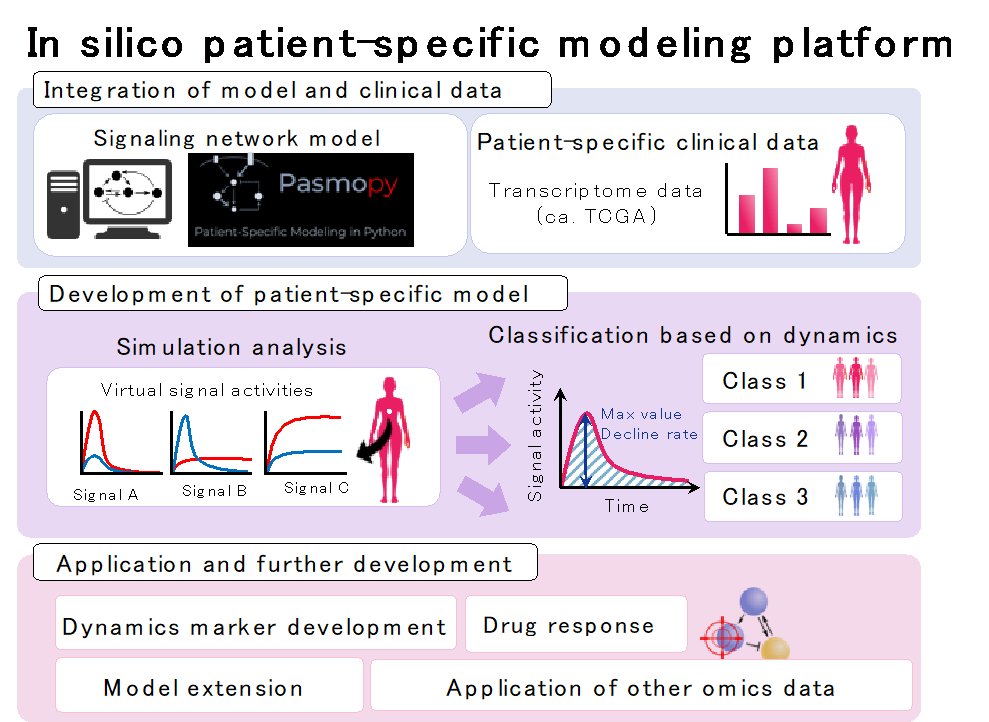
Based on the above-mentioned BioMASS, we are constructing an in silico patient-specific model (virtual patient) that combines mathematical modeling and patient clinical data. This makes it possible to build a personalized mathematical model and perform simulation analysis based on the genetic characteristics of each patient. In addition, from the simulation results, subtype classification, prognosis classification, and drug susceptibility analysis become possible. So far, it has become possible to classify the prognosis of triple-negative breast cancer and predict drug using the transcriptome data obtained from TCGA (The Cancer Genome Atlas). This approach is also applicable to other cancers. This patient-specific model package PasmoPy (Patient-Specific Modeling in Python) can be downloaded from https://pepy.tech/project/pasmopy.
Laboratory of Cell Network Regulation
| Website | http://www.protein.osaka-u.ac.jp/cell_systems/pg18.html |
|---|---|
| mokada*protein.osaka-u.ac.jp (replace * by @) |
